Documentation Index
Fetch the complete documentation index at: https://docs.boltz.bio/llms.txt
Use this file to discover all available pages before exploring further.
Overview
Targets are the protein structures you want to design binders against. You can define targets by entering sequences or uploading existing structure files.Creating a New Target
- From your project dashboard, click ”+ Add target” in the Targets section.
- Enter a Target Name (required).
-
Choose how to define your target:
- Define Sequences
- Upload Structure
Click ”+ Polymer” then “Protein from Sequence”Paste or enter your protein sequence (maximum 1,300 residues)Click “Import”After submitting, Boltz will predict the structure automatically - Click “Continue” to proceed to target initialization.
Target Initialization
After creating a target, you’ll go through a 4-step initialization workflow:Step 1: Verify Unbound Structure
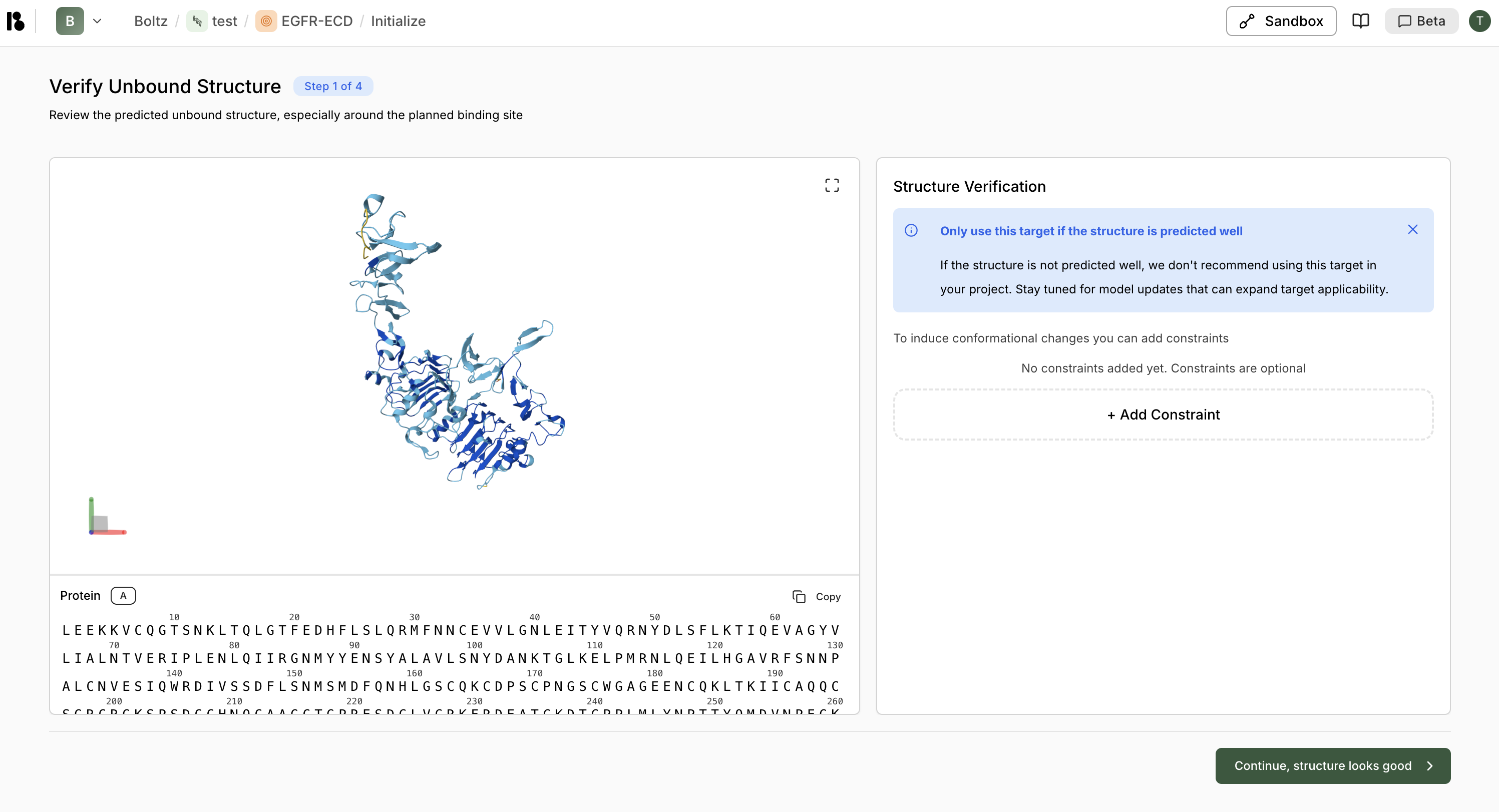
Step 2: Select Crop
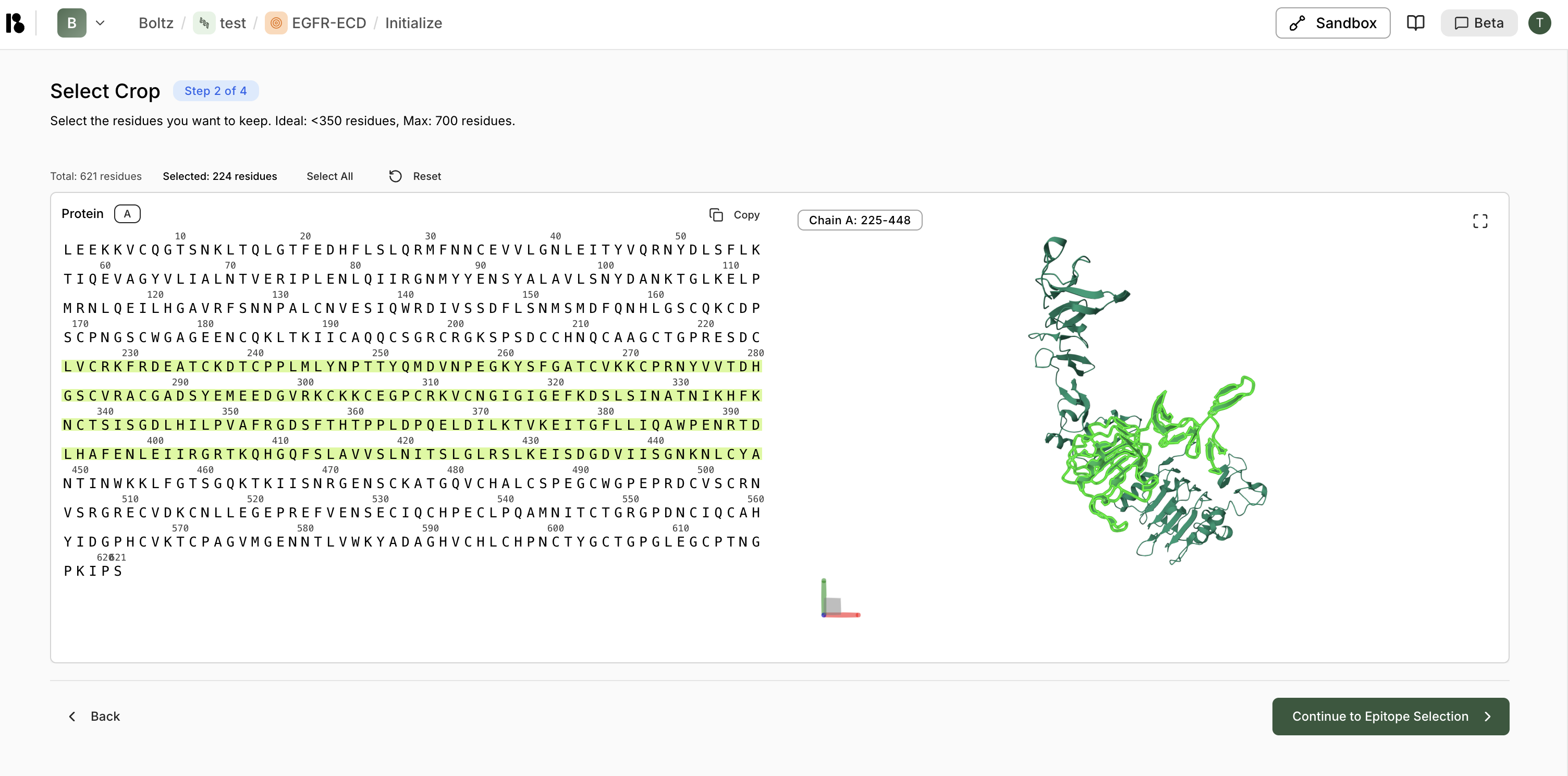
- Use the sequence viewer to select residue ranges
- The 3D structure updates in real-time to show the cropped region
- Click “Select All” to include all residues, or manually select ranges
Step 3: Select Epitope (Optional)
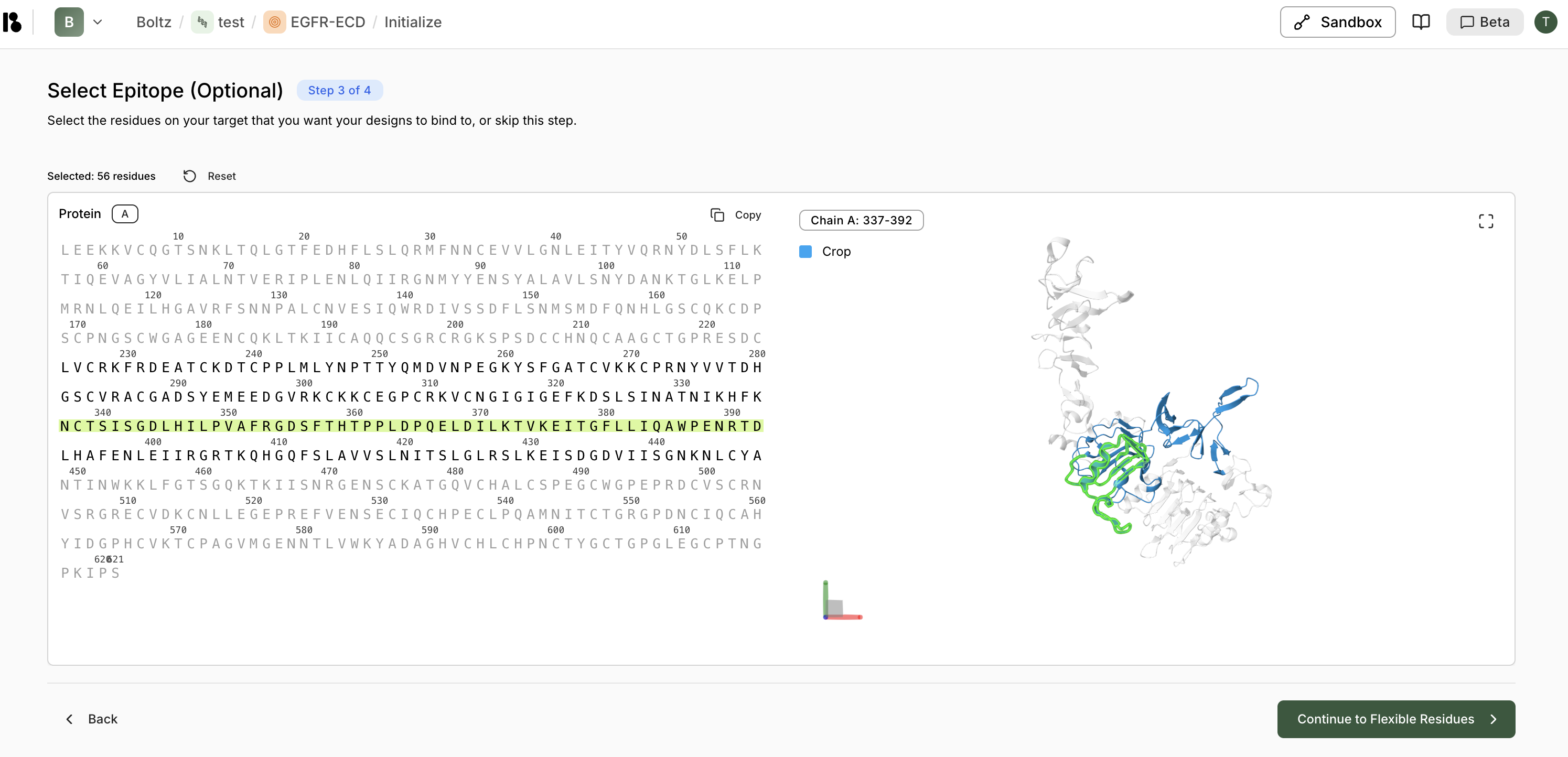
- This step is optional—skip if you don’t want to specify an epitope
- Selected residues are highlighted in green in both sequence and 3D views
- Click “Reset” to clear selections
Step 4: Select Flexible Residues (Optional)
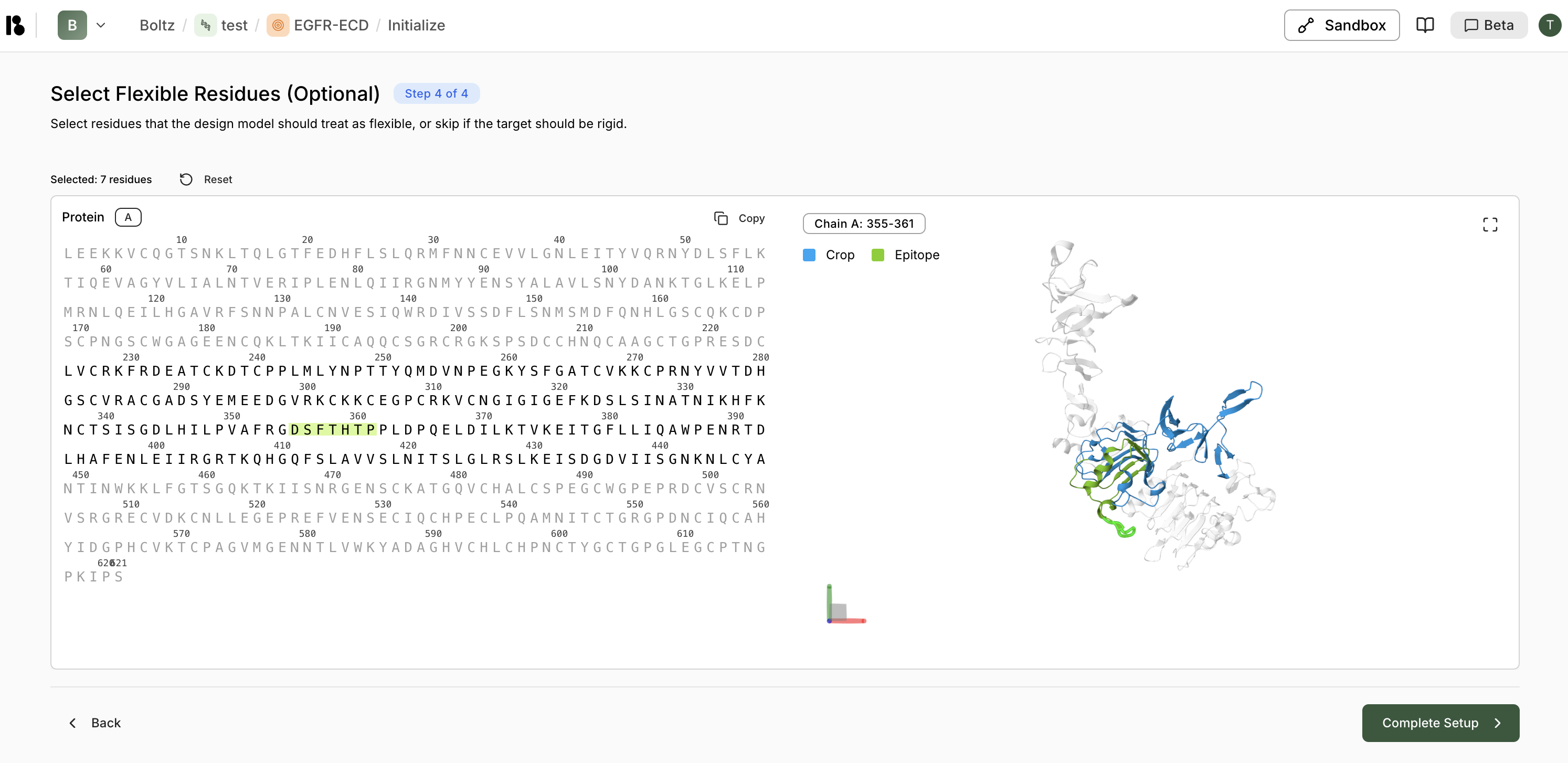
- This step is optional—skip if the target should be treated as rigid
- Flexible residues allow for conformational changes during binding
- Selected residues are highlighted in green
After Target Creation
Once your target is created, you’ll see options to:- Discover New Binders - Create a binder specification for your virtual screen
- Upload candidates from CSV - Evaluate known candidates using AI
- Skip for now - Add the binder specification later