To begin designing binders, add a target protein/receptor via the + Add Target function in the Targets section.Documentation Index
Fetch the complete documentation index at: https://docs.boltz.bio/llms.txt
Use this file to discover all available pages before exploring further.
Adding a Target
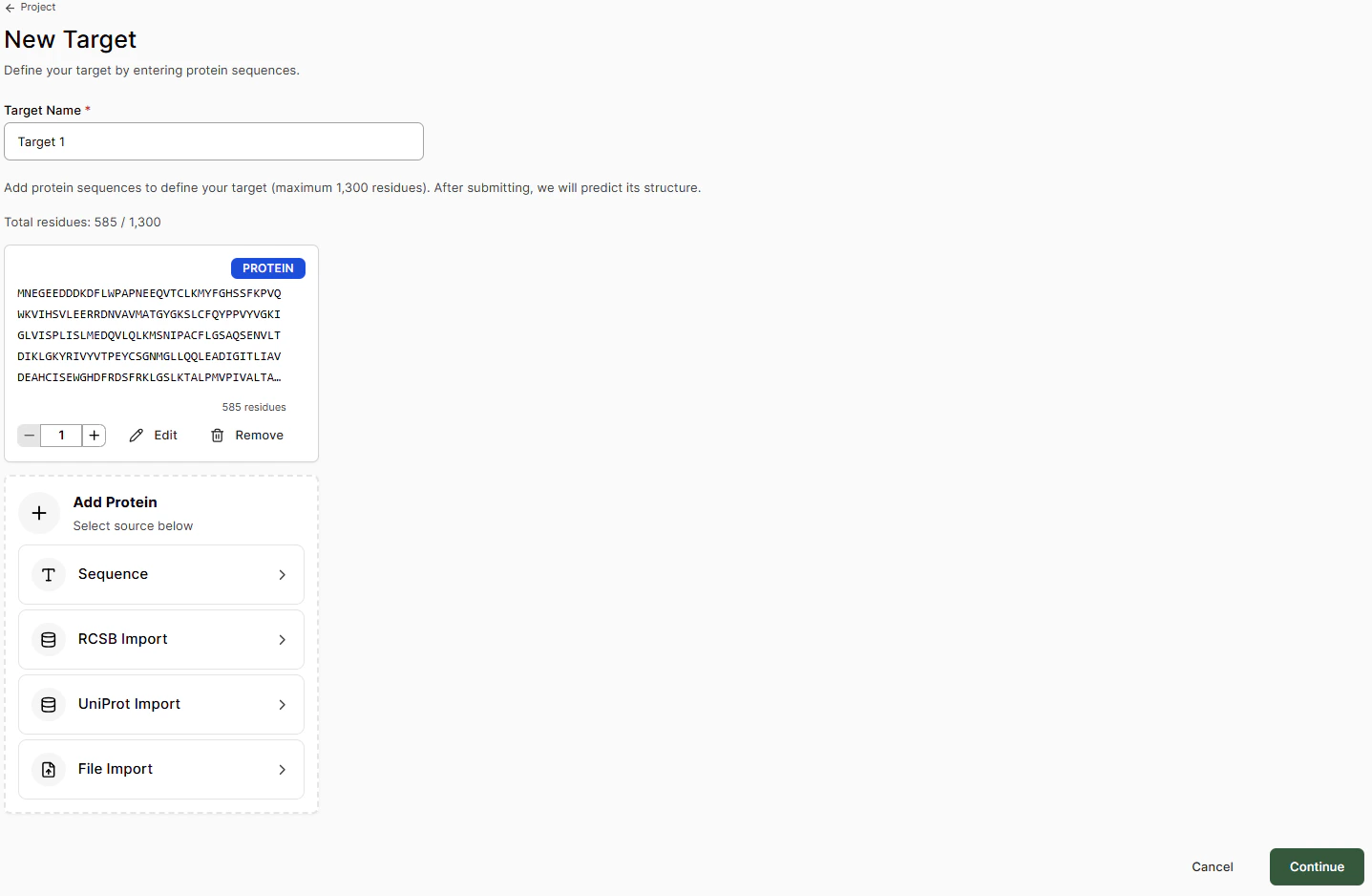
| Import Method | Description |
|---|---|
| Sequence | Paste the FASTA sequence and select any post-translational mutations or modifications |
| RCSB Import | Add sequence via the RCSB-PDB ID code of a deposited crystal structure |
| UniProt Import | Add sequence via the UniProt code |
| File Import | Add sequence from a .cif structure file |
Targets are not restricted to the main receptor of interest in a project. Additional off-target receptors, mutants, homologs, etc., can be added within a Design Project for Boltz structure and binding affinity assessment.
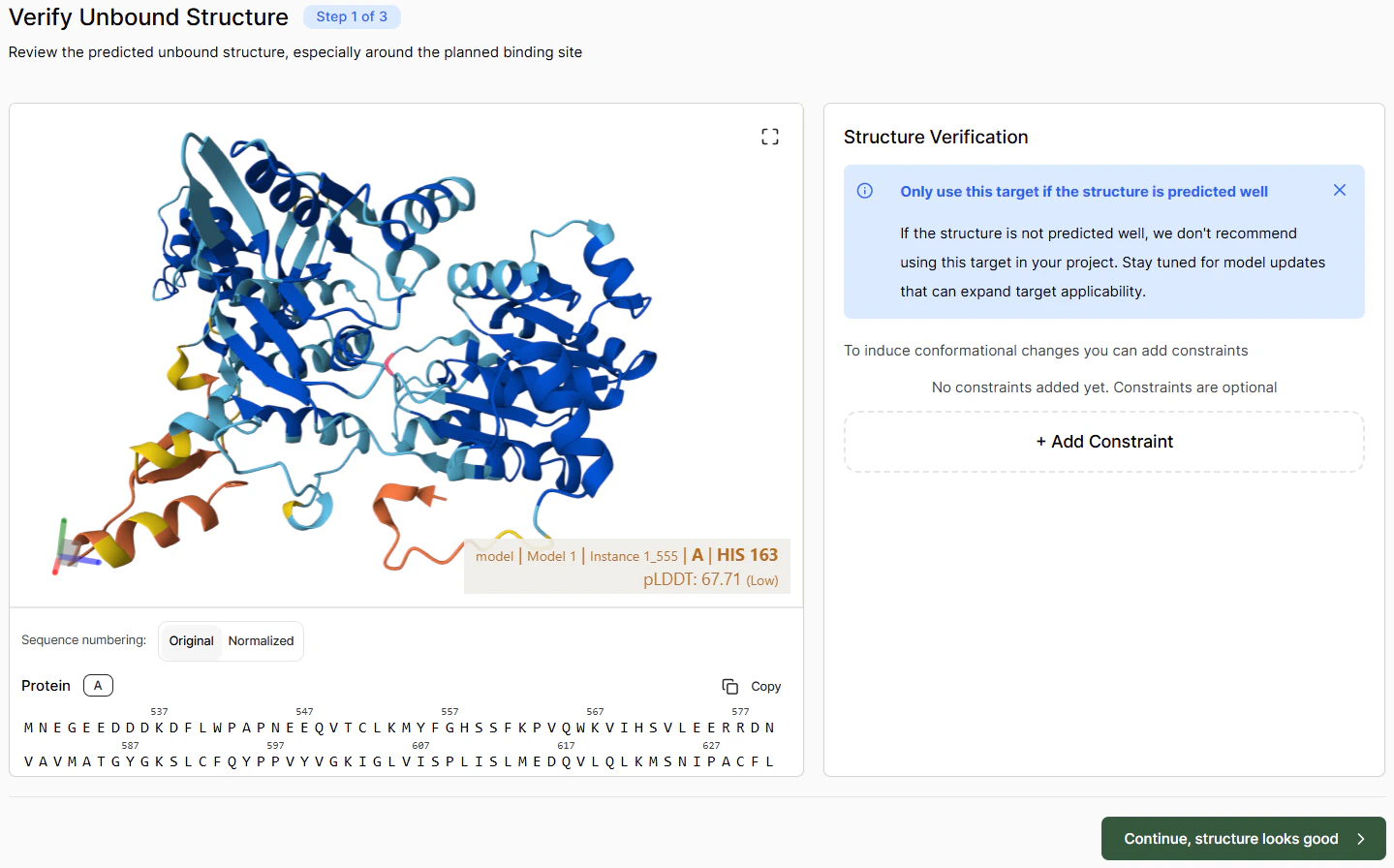
Verifying the Apo Structure
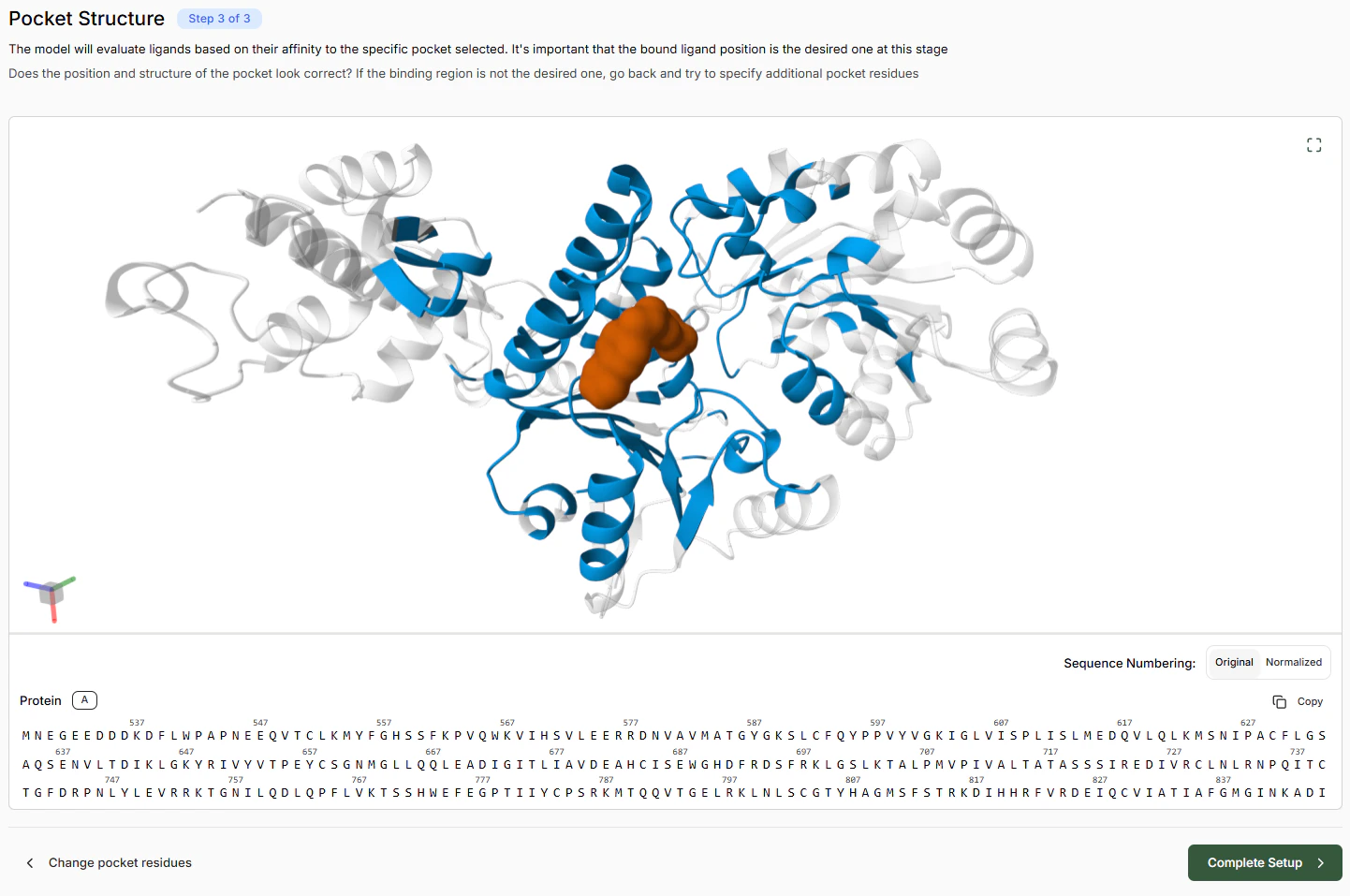
Verify that the unbound structure looks roughly correct. If there are particular issues, you can try solving them by adding constraints. Once satisfied with the apo pose, click Continue.Defining the Binding Pocket
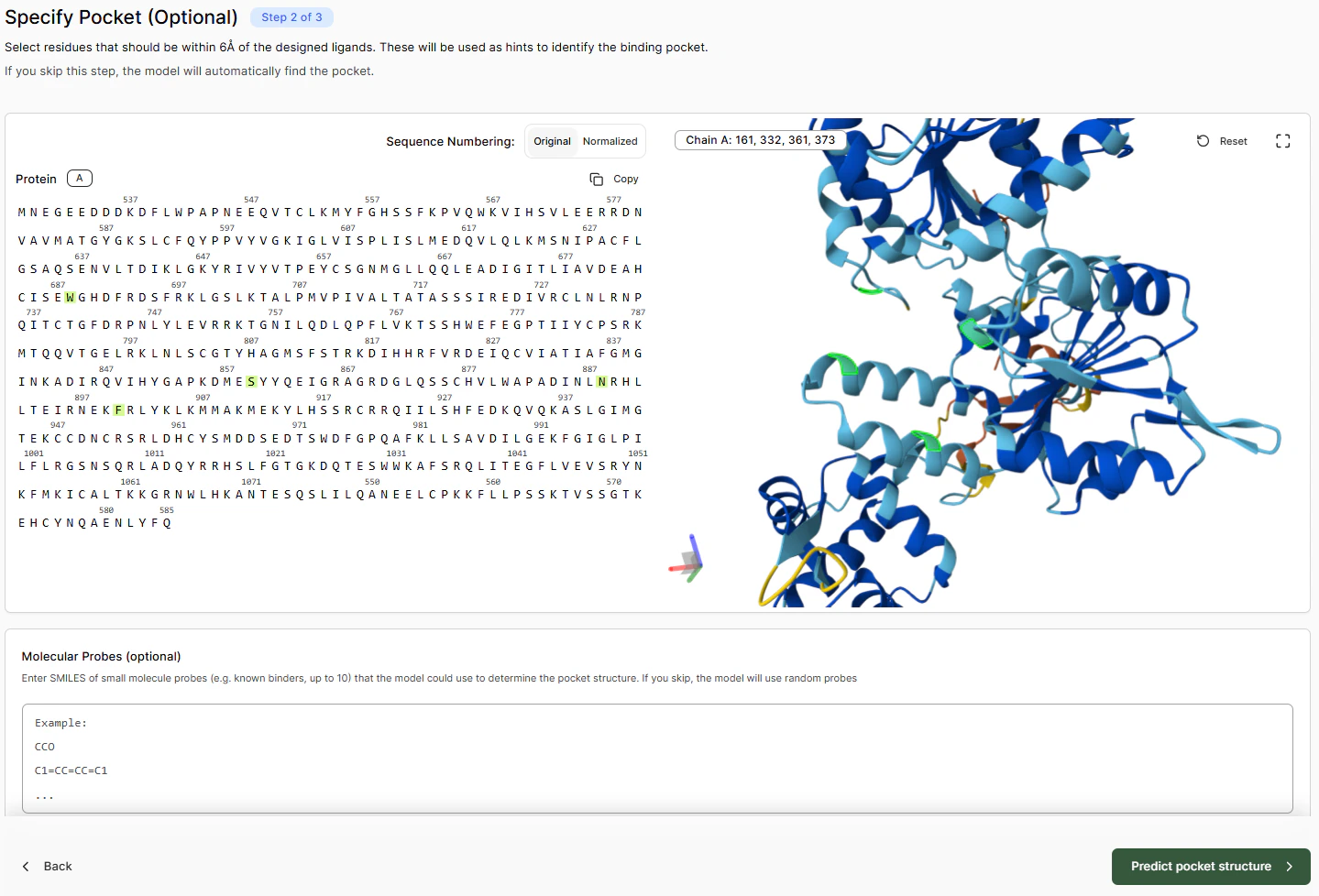
To map out a binding pocket, either:- Select residues in the left-hand sequence panel, or
- Select them directly in the 3D viewer
Deleting Targets
To avoid conflicts, currently Targets that have completed setup cannot be deleted. Targets in progress can be deleted be selecting from the three dot menu in the Target list.